Webinars are powerful lead generation tools. This webinar we hosted for Cantata Biosciences brought in hundreds of new leads. Watch this clip of Dr. Giacomo Cavalli discuss how Polycomb proteins are associated with gene regulation to see how our life science webinars engage the audience.
Webinars are powerful lead generation tools. This webinar we hosted for Cantata Biosciences brought in hundreds of new leads. Watch this clip of Dr. Giacomo Cavalli discuss how Polycomb proteins are associated with gene regulation to see how our life science webinars engage the audience.

With the right marketing strategy and execution, webinars can be a powerful lead generation tool. Samba scientific offers complete turn-key promotional campaigns that drive lead generation before and after the event. Our comprehensive promotional campaigns result in attendance rates that frequently exceed those of our much larger competitors.
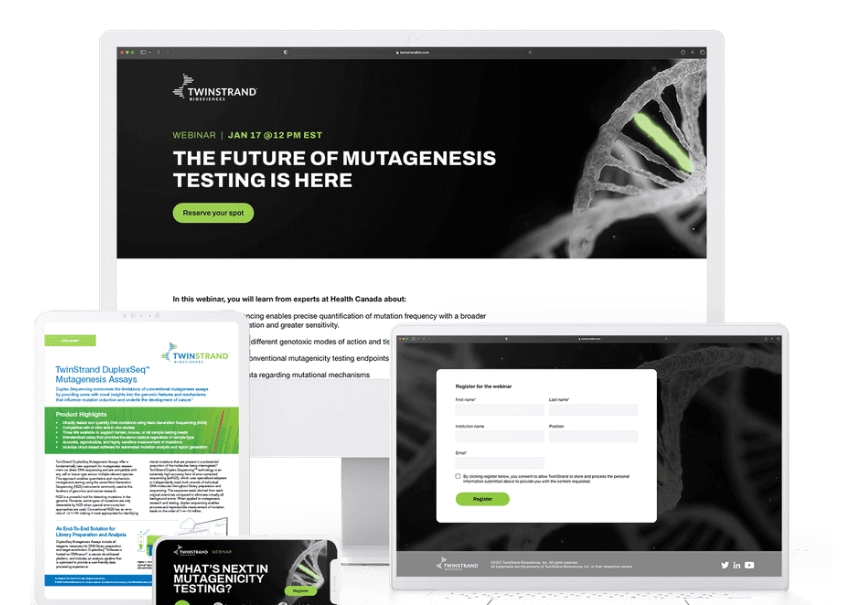
With the right marketing strategy and execution, webinars can be a powerful lead generation tool. Samba scientific offers complete turn-key promotional campaigns that drive lead generation before and after the event. Our comprehensive promotional campaigns result in attendance rates that frequently exceed those of our much larger competitors.
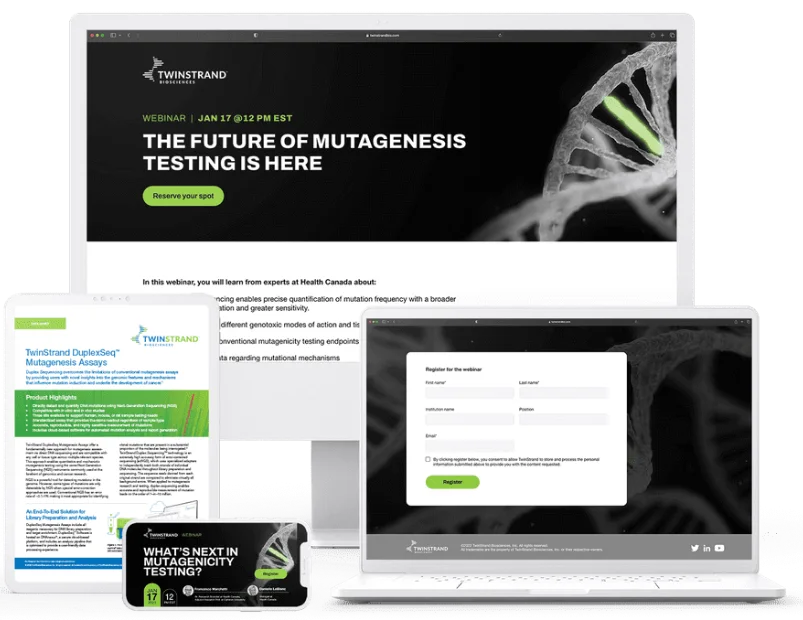
Drive interest and anticipation with promotion across channels, maximizing attendance and capturing leads before your webinar even airs. Our full promotional campaign package includes:
Choose the webinar format to meet your needs, delighting and engaging your target audience with a superior webinar experience. Need a host? No problem! One of our scientific staff members will be available to introduce your speaker and moderate your Q&A session so you can sit back and enjoy the event.
Make your webinar the lead generation gift that keeps on giving, with sophisticated channel deployment, content marketing, and ongoing campaign support. When your event is over, we provide a report with complete information about your new leads. Our post-promotional plan includes:
Now that you have new leads, it’s time to develop nurturing workflows to qualify them for hand-off to your sales team. After your webinar, you can nurture leads with additional webinar-based content:
Get the latest tips, tricks, and news


Get the latest tips, tricks, and news


Copyright © 2024 Samba Scientific. All rights reserved.